- 1D PSYCHE
- 1D TSE-PSYCHE
- 1D Zangger-Sterk
- 1D Selective TOCSY-PSYCHE
- 1D SAPPHIRE
- 1D semi-real-time pure shift
- 2DJ-ZQS-PSYCHE
- 2D F1-PSYCHE-TOCSY
- 2D rtPS-gHSQC
- GEMSTONE
- GEMSTONE-CLIP-COSY
- GEMSTONE-NOESY
- GEMSTONE-ROESY
- GEMSTONE-TOCSY
- PSYCHE-iDOSY
- DOSY Oneshot
- Oneshot-iDISPEL
- 1D FESTA
- REST
- PUREST
- SCALPEL
- PROJECT
- DISPEL
- ODYSSEUS
- DOSY Oneshot (3ax)
- 3D Oneshot-HSQC
- 19F DOSY Oneshot45
- Pure Shift DOSY Oneshot
- 2D Constant-Time nQF-COSY
- PE WATERGATE (solvent suppression)
- 2D Zangger-Sterk NOESY
SCALPEL
Summary:
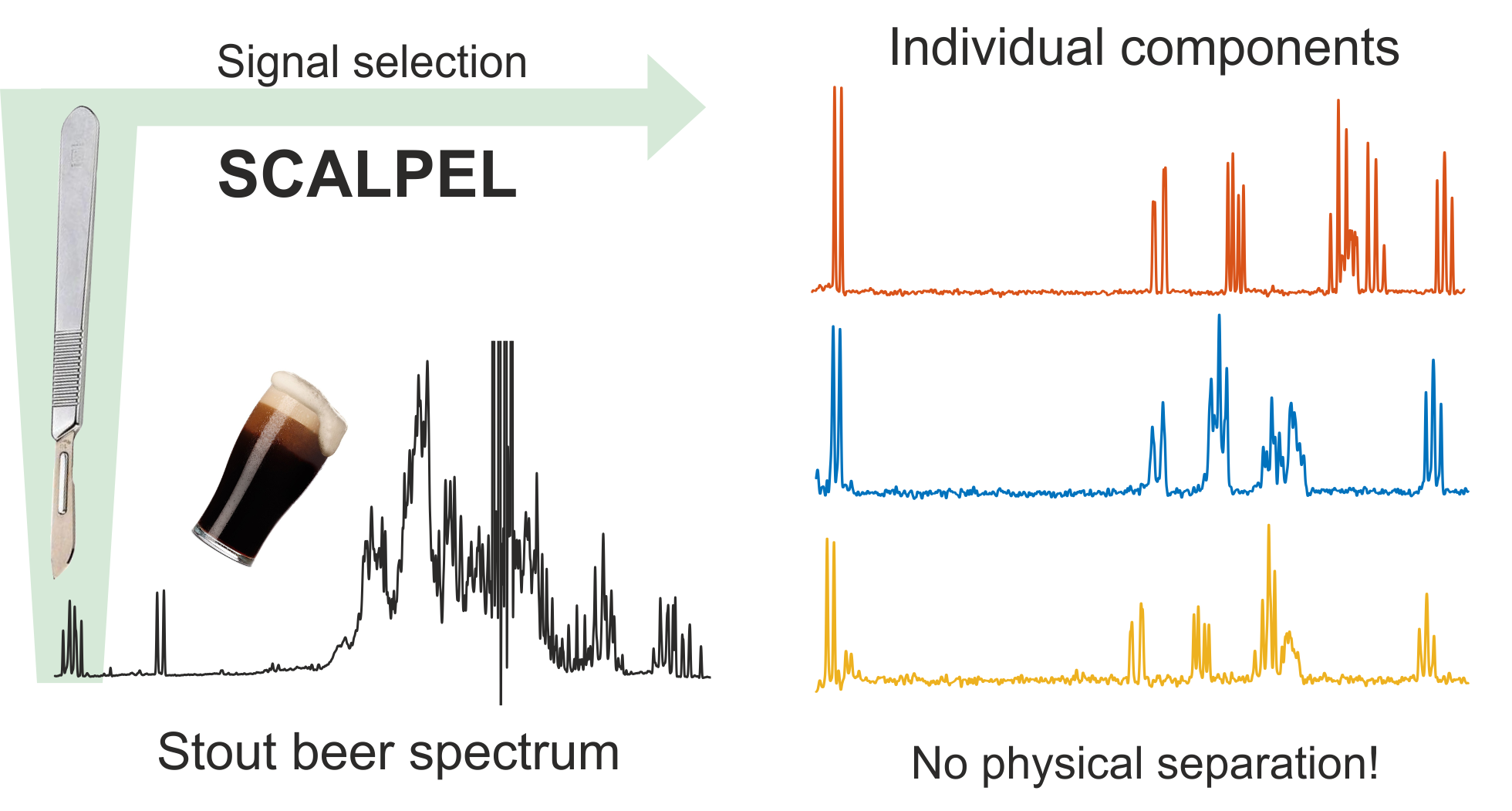
SCALPEL (Spectral Component Acquisition by Localized PARAFAC Extraction of Linear components, SCALPEL) is a new family of NMR experiments that allows the efficient, practical and non-destructive analysis of complex mixtures. This new class of NMR experiment is based on dissecting the spectrum rather than the sample, using pulse sequences tailored to generate data suitable for powerful tensor decomposition methods to allow highly complex spectra to be analyzed stepwise, one small section at a time.
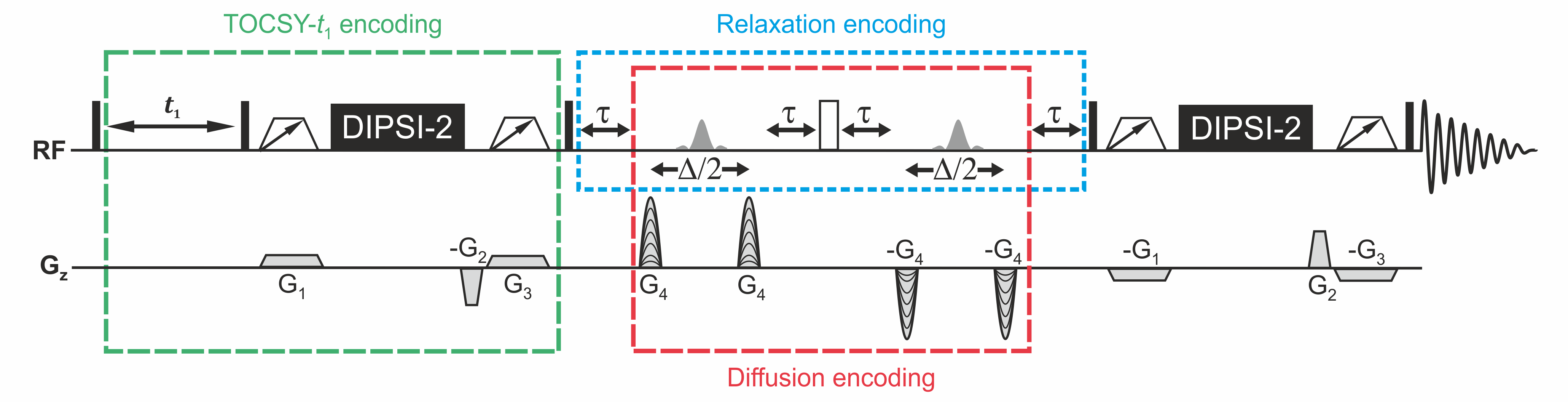
Pulse sequence diagram:
A prototypical pulse sequence for SCALPEL experiments using TOCSY-t1 evolution encoding (outlined in green) and either diffusion (outlined in red) or T2 relaxation (outlined in blue) encoding is shown below. Only a small number of values are used for the evolution time t1, chosen for efficient sampling of differences in signal evolution, rather than the large number of equal increments that would be used in a 2D experiment. For diffusion encoding, the delays τ are kept constant and gradient strengths G4 are incremented, while for relaxation encoding gradient strengths G4 are kept constant and delays τ are incremented.
Downloads:
Bruker pulse program:
SCALPEL with TOCSY-t1, diffusion and T2 relaxation (2022 - updated syntax) (right click and chose 'save ... as')
SCALPEL with TOCSY-t1 and diffusion (right click and chose 'save ... as')
SCALPEL with TOCSY-t1 and T2 relaxation (right click and chose 'save ... as')
Experimental data:
Stout beer
Processing:
The processing of SCALPEL experimental data is straightforward using the free and open source software package GNAT.
Reference:
Dal Poggetto, G.; Castañar, L.; Adams, R. W.; Morris, G. A.; Nilsson, M.; J. Am. Chem. Soc. 2019, 141, 5766-5771.
DOI: 10.1021/jacs.8b13290
Please note that copyright restrictions prevent us from placing the pdf files on our website, however, this article is open access and the pdf can be downloaded from ACS Publications.
